Unix shell pipeline: Making it multi-lane, Avoiding stop-n-go
This is a CS technical post related to using unix command-line. Just an early warning so that you can walk away, saving your precious browsing time! :). BTW I am aware of some poor formatting issues.. please bear with it..
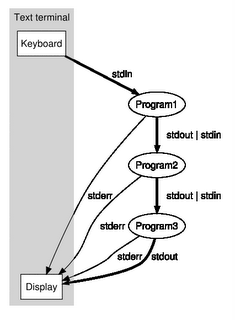
Shell pipeline is the concept of attaching the stdout of a process to the stdin of another process. See wikipedia for more. This is one of those beautiful unix concepts that stood the test of time. It's simple and extremely effective. It's a shame that modern GUI designers didn't even think of a way of supporting the pipeline concept.. anyway I guess that's a different topic altogether.
Take the simple unix pipeline
$ ls | wc
It prints the number of files in the current directory (for the purist, the number of elements.. )
Recently I was looking at a way to search for a string in a directory tree. The earliest grep didn't do recursive grep. I am sure the newer ones sport the -r flag to do that. They even taken patterns to include/exclude directories.
I started off with the very basic, home grown rgrep.
karthikg@xyz :>cat ~/bin/rgrep
find . -type f -a -exec grep -l $1 {} \;
You run it as 'rgrep needle' to search 'needle' in the whole directory tree.
I need to do a search on a source code tree containing about 100k files ( that's just text files not counting binary files). When I issued the command from the commandline, it took forever to complete. And thus began the optimization process.
The very first step is the well-known trick of using xargs to avoid the exec in find. With -exec of find, we are forking a new process for every element, as find descends the tree. And this is very expensive. With xargs, it tries to feed as much files as possible to each created grep process.
find . -type f -print | xargs grep $pattern
Even this took forever; I started to move away from solving the generic case by exploiting known special properties. Some points to note 1. Almost always I'm interested only in text files. Thus using 'file' command, I can quickly discard binary files before they reach grep. 2. The directory tree has lots of object/executable file directories, which can be skipped. This is a big gain since 'find' need not descend the whole subtree.
Thus the next solution became
find . ( -path *obj-ppc -o -path *lib-ppc -o -path *bin-ppc -o -path *\.CC -o
-path */\.T* ) -prune -o -type f -print |
xargs file -- | grep text | cut -d: -f1 |
xargs grep $pattern
Please note shell quoting of special chars (*, .) is not shown above (that's
one other irritating thing.. can't we somehow tell the shell not to do
that??)
Here's what is happening:
- In find, we try as much as possible to skip whole sub-directories. -prune helps us there. I had -regex.. but felt it's too big a gun to use; -path must be light weight and for my needs it does the job.
- Then -type f helps to cut down the elements to regular files
- The second process in the pipeline, gets the file type, the following two, grabbing the text files and the cut gets the file name alone
- The last grep does the heavy weight string searching
I thought the above must run pretty fast. But to my surprise, it was crawling and looked like it will never complete.
I was running on a fairly powerful dual CPU linux machine and a top showed lots of free CPU. So why isn't it completing soon? Where are the bottle-necks?
My first guess is that grep is taking too much time (the last process). I was wondering if I can parallelize the search.. why can't I split the input file-list (of 100k entries) into say 4 chunks and feed to four different grep processes. These 4 can run in parallel and must finish sooner. Here is when I thought of a multi-lane shell pipeline.
Let's take this syntax
p1 | p2 |{number} p3
e.g. find .. | ... |4 xargs grep $pattern
Here I'm telling the shell to use 4 processes to do the last grep work. It must split the input into 4 parts and send them to each of the four xargs. Of course none of today's shell can do this. I ended up writing a python version of the above (see end). I'll discuss more on some of the findings in a later post.
Going back to the original slowness, looks like the good old shell pipeline is causing lots of stop-n-go among the processes. The find can't run in full speed as it fills its output buffer and gets blocked before the next process in the pipeline can read its stdin. Thus even with CPU usage so less, chaining lots of processes didn't help in the throughput. (Later using -fprint of find, I realized the find completed at a fraction, less than 10%, of its time with -print and pipeline)
My multi-lane version performed marginally better.. I make the "file ..cut" part use 4 threads and also the final grep part use another 4 threads.. still the response was sluggish. The bottle-neck seemed that the stop-n-go introduced by connecting processes. I experimented with using temporary files for the output.. and the speed gain was astounding. The whole process got done in a matter of seconds (about 40s) where-as the pipeline one took a whole 14 minutes to complete.
It would be good to have a shell feature where you use intermediate files to connect the processes, rather than directly feeding stdout to stdin. In the early days, avoiding large intermediate data would have appeared a welcoming solution; but today with vast amounts of memory and fast file-systems with large file caches, it makes more sense to run each processe at its full speed, rather than do this stop-n-go due to intermediate buffer issues.
Thus in conclusion I see that the real bottle-neck is not the load of the system, it's the IPC buffering issues introduced by the pipeline. So when high performance is needed, try using temporary files for stdout and stdin, which give enables each process to run at it's top speed. Even though, you are using the disk filesystem, speed gains are significant.
--- multilane_pipeline.py ----
from threading import Thread
# a small worker thread to read out the stdout of a child process
# This is used for proceses which generate lots of output as well as
# take in large stdin
# To solve the classic pipeline deadlock -- ie child can't proceed
# unless someone removes its output; but that someone can't wait to
# push in all the stdin into the child.
# So once a child is forked, before shoving all data into it's stdin,
# make a worker thread using this class. This worker thread will
# remove (periodically..as and when child outputs) child's stdout, so
# that child can continue to make progress
#
# Use only on targets where we push in large stdin *and* target generates
# large stdout. Else not needed... set flag 'is_thread_collect' if you
# anticipate this scenario
#
class stdout_reader (Thread):
def __init__ (self,fd):
Thread.__init__(self)
self.fd = fd # while file object to collect from
self.data_read = [] # the data collected
def run(self): # when giving 'go', pull out all data
self.data_read = self.fd.readlines()
# command processor
# A thread which will run a unix command and collects its output
# It takes in a list containing the whole stdin (thus different from
# standard module commands)
#
class command_processor (Thread):
def __init__ (self, cmd, stdin_list):
Thread.__init__(self)
self.cmd = cmd # Unix cmd.. can be sequence..sent to Popen
self.stdin_sent = stdin_list # What stdin to feed to cmd
self.stdout_rcvd = [] # What output collected from cmd's o/p
self.status = 0 # status of cmd execution
self.status_is_signaled = 0 # status of cmd execution ..is signal?
def run(self):
if options.debug > 0:
print "Starting thread for cmd: %s" % self.cmd
run = popen2.Popen3(self.cmd)
# set up another thread to start reading the stdout ...
reader_thread = stdout_reader(run.fromchild)
reader_thread.start()
# Feed to cmd's stdin
run.tochild.writelines(self.stdin_sent) # push input into child
run.tochild.close()
reader_thread.join() # wait for the worker thread to finish
self.stdout_rcvd = reader_thread.data_read
result = run.wait() # block wait until cmd is finished.
self.status = os.WEXITSTATUS(result)
self.status_is_signaled = os.WIFSIGNALED(result)
#
# Run cmd using given number of threads
# return output and status
#
def execute_child_threads (cmd, stdin, target):
thread_str = target_db[target]['threads']
# Do any $thread to actual value user gave on '-a -t'
num_threads = int(substitute_target_args(target, thread_str))
stdin_size = len(stdin)
if stdin_size < num_threads:
num_threads = 1
work_slice = stdin_size / num_threads; # each thread's work size
workers = []
# We are going to chop stdin into equal sized chunks for each thread.
for i in range(num_threads):
start_index = i * work_slice;
end_index = start_index + work_slice;
if i == num_threads - 1: # if last worker...take all remaining
end_index = stdin_size;
this_stdin = stdin[start_index:end_index] # this thread's in data
cp = command_processor(cmd, this_stdin)
workers.append(cp);
cp.start() # fire up the thread to do the data munching
output = []
for worker in workers:
worker.join() # wait for each worker to finish
output += worker.stdout_rcvd # collect the output it generated
#TODO. Handle each thread's return status...
return (output, 0, 0)


1 Comments:
interesting. thanks.
Post a Comment
<< Home